General vision/ Big picture
We are interested in vesicular trafficking, the signal processing at the level of single cells, and in the context of intercellular communications in development and diseases. Our general philosophy is to go beyond the generic ‘X entity regulates Y process’, and understand the precise events and mechanisms in place and bring forward concepts that are specific.
We use generic concepts in physics, to frame concepts and mechanisms that go beyond the lack of clarity that is inherently associated with describing a protein’s activity as ‘regulates’. Please read this article that also emphasizes our view. While many-component regulatory circuits are present in biology and help classify the plethora of processes and describe the network topology, here, our aim is to identify the precise mechanisms and processes that will help us define well-detailed concepts, scaling from molecular structure, resulting in interactions including non-linear interactions with rich substrates like a membrane, to macroscopic phenomenology relevant to a process inside the cell, and hopefully extending to tissue level phenomena.
Seemingly chaotic, do molecular systems extract order?
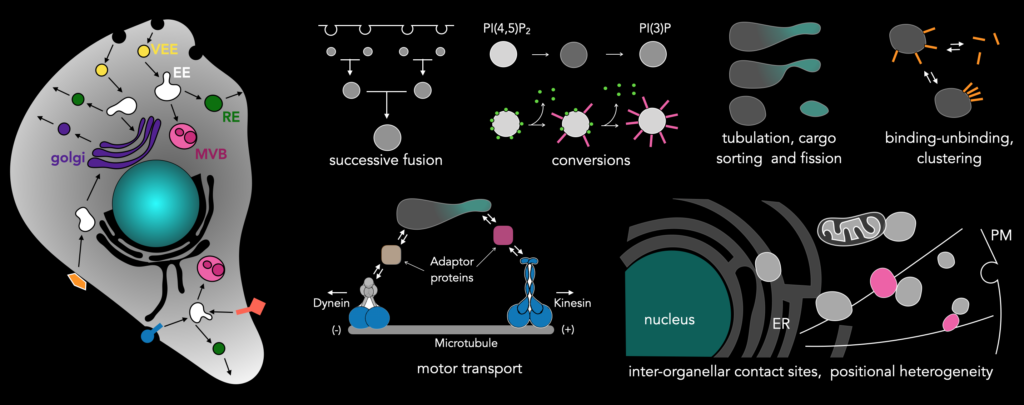
We are currently focusing on the endosomal network, the intracellular transport system of the cell. Individual processes that build up the endosomal network are intrinsically stochastic. These processes are endosomal fusions, cargo sorting, tubulations and fission, binding of adaptor proteins and downstream motor proteins, endosomal contacts with other organelles like the ER or other endosomes, biochemical interactions leading to endosomal conversions. All the processes that constitute the endosomal network happen seemingly stochastic and are interlinked. We hypothesize that the molecular-network architecture of protein involved in the endosomal network encodes various mechanisms by which temporal precision is maintained, or in other words, extracts order from chaos. Our approach of using rapid whole-cell imaging using LLSM allows capturing all events pertaining to proteins of interest that are relevant to endosomal processes. Even though the events are stochastic, they will be captured when imaged at high temporal resolution in the entirety of the cell, which can later be visualized and analyzed.
Along these ideas, a concept we have been exploring in the lab is that of time-keeping of processes in the cells. What molecular mechanisms set the rate of endosomal maturation, a process defined by the localization of specific ‘marker proteins’ that defines the local biochemistry. What is the first passage time for the receptor arrival? What are the implications in interpreting a signal (e.g. a ligand pulse in time) for the cell? What mechanisms operate to distinguish distinct ligand concentrations?
We hypothesize that the endosomal trafficking system is the main logistical machinery for such ‘cognitive’ processes by a single cell, and hence we strive to understand the endosomal network on its own as well as in the context of signaling receptors. Please read our review for more on this.
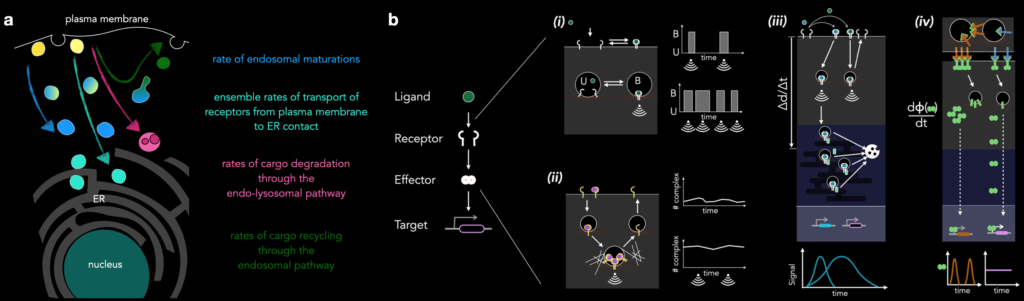
Major focus of the lab:
